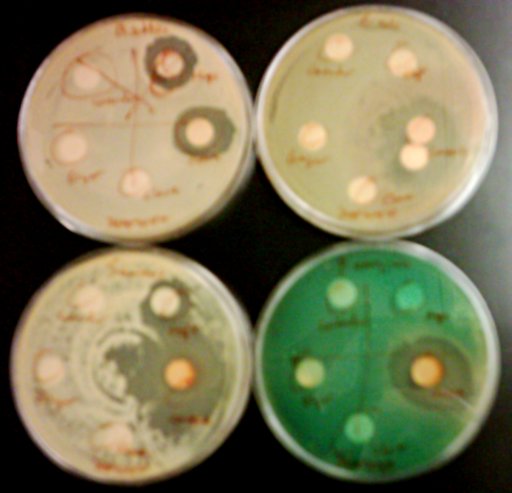
In a typical student microbiology lab, it seems whenever you get your hands on some bacteria the first thing you do is check to see if it’s “Gram-positive” or “Gram-negative”. But does this old Victorian-era test still mean anything useful in the context of modern bacterial taxonomy?
It would seem that it actually does. In my admittedly limited experience, an un-ambiguous Gram-positive result using the standard procedure nearly always indicates bacteria in the phylum firmicutes, sometimes also referred to as the “low G+C gram positives”.
The “low G+C” part has to do with the chemical characteristics of the DNA. If you’re already familiar with DNA’s structure then you probably already know what this means, but for everyone else, in brief:
DNA is made of strings of four different chemical “bases” chained together. The bases are “Adenine”, “Guanine”, “Cytosine”, and “Thymine” (abbreviated as A, G, C, and T). These bases make up a sort of chemical “alphabet”, which encode how to make various proteins. These chemical bases each have an opposite base that they are attracted to – Adenine to Thymine, Cytosine to Guanine, so DNA’s strings end up matched with an “opposite” string, which together form the easily recognized “double helix” shape of the overall DNA molecule. The relevance here is that the attraction between the Guanine and Cytosine (“G+C”) is stronger than the attraction between Adenine and Thymine, so you can estimate relatively how much Guanine and Cytosine is in an organism’s DNA by how tightly the two strands of DNA stick together.
And, yes, there is a “high G+C” gram-positive group, with a larger proportion of the G and C bases as compared to the A and T. That’s the “Actinobacteria”, which includes the “acid-fast” Mycobacterium group. But that’s a topic for another post.
The firmicutes, with the exception of one group that I know of, all have a distinctive, simple outer structure. It’s something like this:
Imagine a water balloon filled with lime Jell-O�. Now, tightly wrap the water balloon with a piece of thick, padded packing blanket. The thick layer of packing blanket is the cell wall. The balloon is the inner cell membrane. The Jell-O� is the cytoplasm. (It’s Lime merely because I like lime Jell-O�.) It might be worth noting that since this group of bacteria relies so much on it’s cell wall, they are more likely to be killed off by the ?-lactam antibiotics (which specifically attack bacterial cell wall generation) than other types of bacteria.
There is one exception to this structure – the class of firmicutes known as the mollicutes. There’s one example of this group that gets mentioned in basic medical-centric microbiology classes: the genus Mycoplasma (as in Mycoplasma pneumoniae). Members of this class have lost their ability to make cell walls entirely.
By contrast, a typical “Gram-negative” cell has a more complicated outer makeup. In brief, start with the same lime Jell-O� balloon, but instead of a thick packing blanket, wrap it with a single thin layer of cloth, and then stuff the whole thing inside of another lime Jell-O� balloon. You can also imagine a bunch of valves stuck through the outer balloon to let stuff in and out, but then you end up imagining the Jell-O� spurting out all over the place and the whole analogy breaks down. If it helps, you can also imagine that if you punched out a section of a gram-negative outer structure with a cookie-cutter, you’d get something kind of like an inverted Oreo� cookie, with easily-dissolved filling on either side of a thin layer of cookie. Anyway, the outer balloon represents the outer cell membrane, the cloth is the (much smaller) cell wall, and the layer of lime Jell-O� between the outer balloon and the cell wall is called the “periplasmic space”.
Interestingly, despite this difference in complexity, the molecular evidence[1,2] seems to indicate that the simpler firmicutes diverged evolutionarily from the more complex “gram-negative” types, not the other way around. The lineage suggested by the 16s gene sequences implies that both types of Gram-positives split off from the Gram-negative type bacteria as a single ancestral group, and some time later the two types diverged into separate groups. If I had to bet, I’d personally put my money on the evolutionary sequence going Gram-negative-type->actinobacteria->firmicutes.
Another characteristic of some (but not all) of the firmicutes is the production of “endospores”. Unlike the spores of molds or Myxobacteria, endospores aren’t reproductive. Instead, they’re a like a lifeboat, or perhaps a metaphorical bomb-shelter in the cells’ also-metaphorical basement. If environmental conditions get unpleasant, the bacteria essentially pull their DNA and a few necessary enzymes into a small, thick, multilayered compartment – the endospore – where they can wait, protected and dormant, until conditions become comfortable again.
The special “Endospore stain[3]” uses a dye called Malachite Green and works somewhat similarly to the Gram and “Acid-fast” stains – the extra-thick spore coats retain the green stain (once it’s been driven in by some extra heat) while the decolorization rinse washes it out of everything else. I still need to try using a microwave oven for the heating step…but that, too, is a topic for another post.
If your microbiology class was typical, when you think of the firmicutes, you may think of little more than “Strep throat”, “Anthrax”, and “MRSA“. If so, though, you’re missing out on the useful ones.
Since Gluttony is my second most favorite “Deadly Sin�”, I tend to think of food-related possibilities here. And, no, I don’t mean Botulism.
Yogurt and Kumiss and Kefir, Sauerkraut and Kimchi, Sourdough bread, Salami, and Belgian Lambic ales all involve (at least in part) growth of various lactic acid producing firmicutes. Mostly members of the Lactobacillus genus, though if you take a look at the labels of some of the “Live and Active Cultures” yogurt you may also spot a close relative of the ‘Strep throat’ bacterium in the list. These kinds of bacteria can be happy members of the “Normal Flora” of the healthy human gut.
Another, more obscure example is “Natto”. A strain of Bacillus subtilis, originally derived from rice straw, is allowed to ferment soybeans. The end result is a pungent mass of beans, covered with gooey slime, and having an odor vaguely resembling old cheese. I’ve actually eaten it, and they aren’t nearly as bad as this makes them sound…though I personally still haven’t acquired a taste for them. You may have seen this stuff if you watch Iron Chef – it was used as the Secret Ingredient in one episode. There does appear to be some real potential for it as a “health food”, though, which you can see if you poke around Google or PubMed searching for “Natto”. Maybe next time you end up in the basic Microbiology lab and you’re given some B.subtilis to look at you can whip out a jar of soybeans to inoculate while you’re at it. (Note that I wouldn’t actually recommend eating the results in this case…)
Comments and corrections, as always, are welcome.
P.S. no affiliation with any of the trademarks mentioned should be inferred here – I just figured the trademarked names would be more recognized than terms like “gelatin” or “sandwich-cookie”…
[1]Sheridan PP, Freeman KH, Brenchley JE: “Estimated minimal divergence times of the major bacterial and archaeal phyla.” Geomicrobiol J 2003, 20:1-14.
[2]Battistuzzi1 FU, Feijao A, Hedges SB: “A genomic timescale of prokaryote evolution: insights into the origin of methanogenesis, phototrophy, and the colonization of land.” BMC Evolutionary Biology 2004, 4:44
[3]Schaeffer AB, Fulton MD: “A simplified method for staining endospores.” Science 1933 77:194
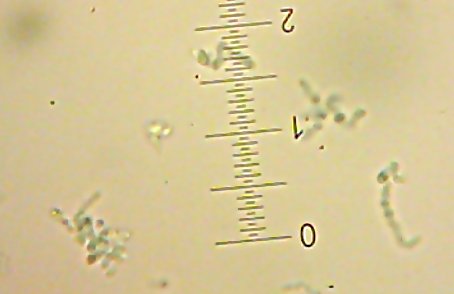
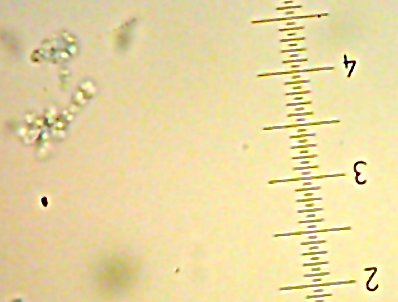
![Sid the [lacto?]bacillus-type-thing](http://www.bigroom.org/images/Sid.jpg)